jackmell
- 1,806
- 54
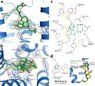
Endorphins in the body act as chemical signals to the morphine-opium receptors (MOR) in the brain and morphine derivatives also react to this receptor. For example alpha-endorphin is a peptide with 10 amino acids and morphine has an aromatic 5-ring non-planer structure.
What set of physical/chemical properties of morphine resemble physical/chemical properties of some part of alpha-endorphin that allows morphine to substitute as endorphin and react with the receptor like endorphin?
Edit: I think I can make the question more precise:
Does the conformational and electrical properties of morphine resemble the same properties of some portion of the endorphin peptide and if so, is that particular region of the endorphin molecule involved with the bonding at the active site of the receptor?
Edit 2: Morphine has a phenanthrene backbone. Phenanthrene is a ring with three fused benzene rings. And so are there other (non-morphine derived) phenanthrene derivatives capable of exerting similar effects as morphine?
What set of physical/chemical properties of morphine resemble physical/chemical properties of some part of alpha-endorphin that allows morphine to substitute as endorphin and react with the receptor like endorphin?
Edit: I think I can make the question more precise:
Does the conformational and electrical properties of morphine resemble the same properties of some portion of the endorphin peptide and if so, is that particular region of the endorphin molecule involved with the bonding at the active site of the receptor?
Edit 2: Morphine has a phenanthrene backbone. Phenanthrene is a ring with three fused benzene rings. And so are there other (non-morphine derived) phenanthrene derivatives capable of exerting similar effects as morphine?
Last edited: