Discussion Overview
The discussion centers on the use of nanopores for DNA sequencing, specifically the role of micron-sized beads in the process. Participants explore the mechanics of how these beads interact with DNA and the implications for sequencing accuracy and methodology.
Discussion Character
- Exploratory
- Technical explanation
- Debate/contested
Main Points Raised
- One participant seeks clarification on how beads assist in nanopore sequencing and the function of optical tweezers in manipulating DNA.
- Another participant explains that while nanopore sequencing does not inherently require DNA to be attached to beads, beads can help slow down the DNA's passage through the pore, allowing for more accurate base identification.
- It is noted that multiple DNA strands are often attached to a bead due to difficulties in creating beads with a single DNA molecule.
- A participant questions whether the DNA is pulled through the nanopore by the bead and if the pulling force varies with each base, contrasting this with the measurement of voltage across the nanopore.
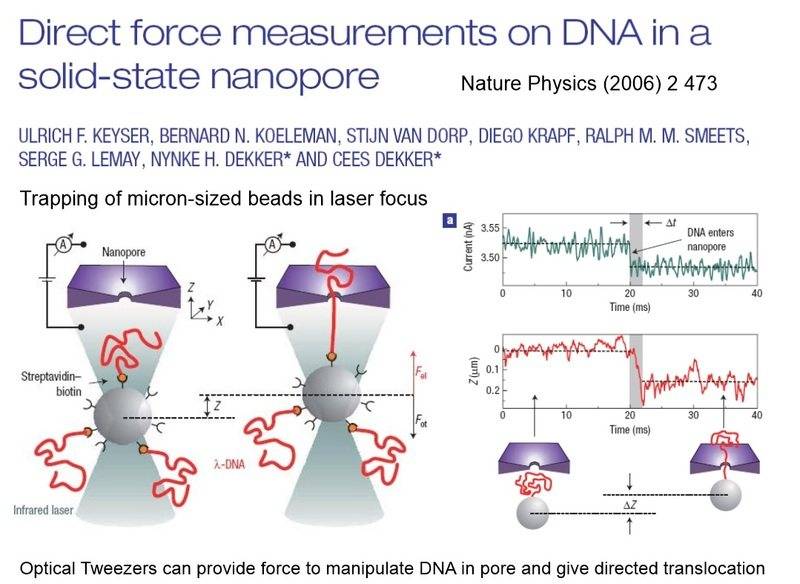
- Reference to a paper is made, indicating that beads are trapped with optical tweezers and brought close to the nanopore, with specific voltage conditions applied to prevent DNA from entering initially.
- There is uncertainty expressed regarding whether the pulling force changes for different DNA bases.
Areas of Agreement / Disagreement
Participants generally agree on the role of beads in slowing down DNA for sequencing, but there is no consensus on the specifics of how the DNA interacts with the beads and nanopore, particularly regarding the measurement of forces and voltages.
Contextual Notes
Some assumptions about the mechanics of DNA movement through the nanopore and the role of optical tweezers remain unresolved. The discussion references a specific paper for further details, but participants have not fully analyzed it.