Hi Sameera!

sameeralord said:
Hello everyone,
I'm very new to microbiology and does not have any background knowledge, so I have some simple questions on identifying different bacteria on agar plates. I have some questions and I have answered them the way I think. Please correct me where nessecary.
1. First of all what do all the lines in that plate mean, and how do you make a culture plate?
The lines are streaks. You inoculate the plate using a loop (a small circular-tipped thing) which has been dipped into a bacterial inoculum (which can either be a broth that's containing living bacteria or a clinical specimen like urine or sputum that contains bacteria) and then use zig-zag motions to layer the bacterial cells onto the agar surface. You zig-zag a few times (forming a few connected "Z" shapes) in one orientation, then change the orientation of the loop so that a fresh edge of it touches the agar. Touching only the last streak of the previous "Z" pattern, streak again at an angle to the previous zig-zags. Do this again once or twice. The key is to be able to get isolated colonies, where each circular mound of growth (called a colony forming unit, or cfu, for short) represents the outgrowth from a single bacterial cell. If there's more than one bacterial morphotype (meaning different looking types of bacterial colonies) which may represent different species of bacteria, you'll be able to tell by inspecting the isolated colonies. The earliest patch where you laid the inoculum down on the agar plate has the heaviest, messiest mixture of growth, and this is called the "confluent part" of the bacterial growth.
To make the agar plate, you either have to do it yourself or buy it commercially. Most modern labs buy the routine stuff commercially, but bigger labs (like mine) retain the capability of pouring their own plates.
To do the latter, you need to follow a "recipe". There are many different types of bacteriologic agar formulations, and you can find the recipes in various reference texts. Probably the commonest type of plate used in a clinical bacteriology lab for primary isolation of bacteria is called a trypticase soy agar (TSA) with 5% sheep blood - which consists of some ingredients plus agar base plus defibrinated sheep blood making 5% by volume of the total. If you're making it yourself, you'll basically melt the agar base down, mix in all the other ingredients and pour carefully into presterilised empty plates to a predetermined depth. Then allow the agar to set, and store at 4 degrees C until they're needed. This is considered too much of a hassle for the common plates, so most labs just buy the stuff, as I said.
Ok so you get the agar medium, and dip a bacterial loop into your sample eg CSF and then streak them on agar plate. So I'm thinking those lines are bacterial streaks.
Yes.
2. Now how do I know there is growth in the culture plate and what is the difference between streaks and circles in the above pic?
Do the circles mean bacterial colonies , then what do the streaks mean?I would like to see an agar plate where there is no bacterial growth, how does that agar plate look like, does it still have original
streaks, or would it just be the agar without any streaks.
Streaks are the lines where your loop contacted the agar. The mounds or "circles", as you refer to them, are the colony forming units I mentioned.
If there's growth, you'll see bacterial colonies (the mounds). If there's no growth, you won't see them. You'll just see the indentations in the agar caused by your streaking.
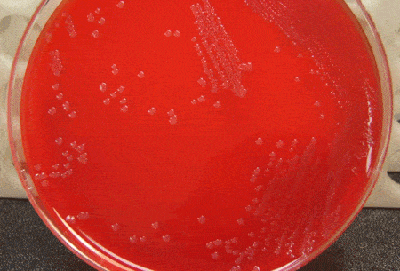
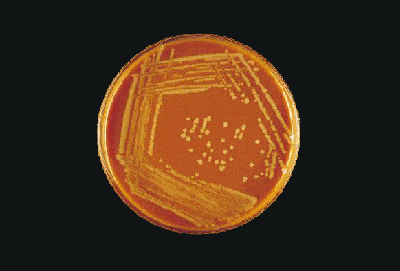
3. Ok this is a picture of proper streaking and growth of H. influenzae on a CAP. Now how do I differentiate this bacteria from other bacteria capable of growing on choclate agar. Also my orginal pic is proper streaking and growth of N. meningitidis on a BAP, same question prevails for this.
Chocolate agar is actually made from lysed (heat-denatured) blood (this is what changes the colour from red to chocolate-brown). This is done to release certain factors called X and V factors into the agar medium to support the growth of certain "fastidious" organisms like H. influenzae.
If you try to grow H. influenzae on just plain old TSA with 5% sheep-blood, you'll find that it won't grow, or it will grow very poorly. To help it grow on this medium, you'll need to include a "streak" of Staphylococcus aureus at right angles to the confluent part of the H. influenzae inoculum. This is colloquially known as a "Staph streak". The Staph streak provides the X and V factors, the former by lysing the blood (Staphylococcus aureus secretes an enzyme called hemolysin that breaks up the blood in the agar plate), the latter factor directly produced by the Staphylococcus organisms. You will observe growth only around the Staph streak (or between the streaks if you did two in parallel), and nowhere else on the plate. This phenomenon is called "satellitism" (like planets satelliting around a sun) and is an identifying feature of H. influenzae and a few other organisms.
Identifying features for H. influenzae:
1) Glistening, gray, non-haemolytic (meaning they don't lyse [split-up] red blood cells, so there's no clearing of the blood on the plate) colonies on chocolate agar plate
2) Satellitism around staph streaks on a TSA sheep blood plate.
3) Growth on an unsupplemented agar plate around a disc with combined X and V factor but not around discs containing only X or V factor singly.
4) Gram-stain: Gram-negative coccobacilli (meaning very short bacilli, that may even appear as round dots).
Other biochemical tests can be done to confirm the identity, or you can use a biochemical kit (like the API NH kit from Biomerieux), or still other methods such as mass-spectrometry or genomic sequencing. The latter (16s rDNA PCR-sequencing) is used only in research settings or, in clinical lab settings, for the "really, really tough to identify by other means" sort of bacteria. Mass-spectrometry used to be mainly a research tool, but it's exploded onto the clinical lab scene, and a lot of labs are routinely using it (a technique called MALDI TOF MS) to identify bacteria grown from clinical samples very, very rapidly (in minutes).
OK, for N. meningitidis:
1) Gray, glistening colonies on chocolate agar (may be somewhat similar to H. influenzae in appearance).
2) Oxidase-positive (a simple biochemical test, but not one that distinguishes it from H. influenzae, since it is also oxidase positive).
3) Gram stain: Gram-negative diplococci - VERY important in distinguishing it from H. influenzae. The cells are arranged in pairs, and each cell in the pair is slightly kidney-shaped.
4) Not X or V-factor dependent (unlike H. influenzae, it can grow on TSA-sheep blood agar without a staph streak to "help" it).
5) Will agglutinate with specific antisera to polysaccharide capsular antigens - this is the basis for "serogrouping" a strain of N. meningitidis to one of A, B, C, W-135 or Y. Note that some strains fail to react to any of these and are described as "ungroupable".
6) Various other biochemical tests can be used to confirm N. meningitidis - the API NH kit should be able to differentiate it from H. influenzae. Within the Neisseria genus itself, certain sugar-hydrolysis tests are often employed to identify N. meningitidis (which should be capable of hydrolysing glucose and maltose, but neither sucrose nor lactose).
Be warned: N. meningitidis (colloquially known as "meningococcus") is a *very* hazardous organism to work with in the laboratory. Unless you're using a specially attenuated (weakened) strain, you shouldn't be playing with it if you're inexperienced. You have to work with cultures and specimens in a biosafety cabinet at all times, and follow all the rules.
Are bacterias able to grow on all agar types or do they have specific ones.
(BTW, it's "bacteria" without the 's'. This is already the plural form, the singular is bacterium).
There are "general purpose" plates like the sheep blood agar and chocolate agar plates. These are often called "unselective" plates because they don't contain factors that specifically suppress certain classes of organisms.
"Selective" plates contain antibiotics or other substances to suppress certain organism classes. E.g. Macconkey agar contains bile salts that suppress many types of gram-positive organisms.
"Differential" plates contain certain indicator dyes that change colour when specific organisms grow on them (mediated by a biochemical reaction). The CHROMagar series is a good example of a differential agar plate that allows the identification of various types of bacterial colonies growing on them simply by colour alone. Macconkey agar itself is also differential (along with being selective), since it contains lactose in the medium, along with an indicator dye, that causes lactose-fermenting colonies like E. coli to turn pink.
If anyone can answer thanks a lot in advance

I'll post more pics depending on the replies.
I hope this has helped. Remember that microbiology is a HUGE field, and I can't do justice over a forum like this. You have to study, study, study and practise, practise, practise.

 I'll post more pics depending on the replies.
I'll post more pics depending on the replies.