jaumzaum
- 433
- 33
Hello guys. I want serious help here.
I'm in first year of medical school and I jut had my second test in Genetics. One question of the test gave us a heredogram and asked us what was the most probable type of genetic disease the family had. There were 2 types of disease that would fit the heredogram. Recessive autosomal and Dominant X-linked.
I put both and explained why both them fit. But a lot of my colleagues put only one of them. He said after that he wanted X-linked, because the heredogram had more trends of X-linked diseases. I just don't bite that.
This is not the way we calculate probability. For example a autosomal disease can happen in 44 out of the 46 chromosomes while the X-linked only in the X chromossome (1.5 in average). Also, what if recessive diseases are more common than dominant. What if most of the people with X-linked diseases are infertile or if most of them dies. All of these reduces the chance of a given heredogram being Dominant X-linked. We can't say something one based on the trends.
The heredogram is the below.
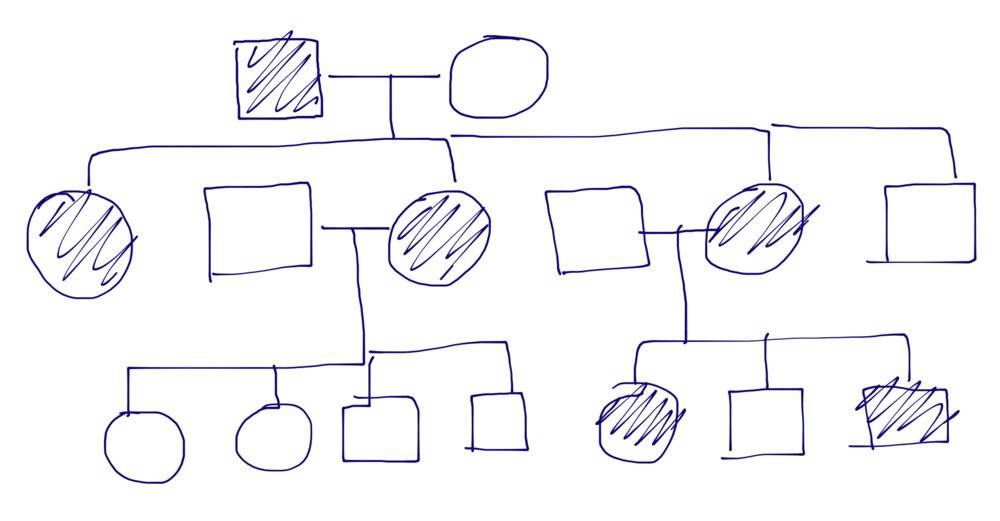
Do you agree with me? I really need some opinions of other people so we can contest the teacher. Otherwise half of the classroom will get a zero.
I'm in first year of medical school and I jut had my second test in Genetics. One question of the test gave us a heredogram and asked us what was the most probable type of genetic disease the family had. There were 2 types of disease that would fit the heredogram. Recessive autosomal and Dominant X-linked.
I put both and explained why both them fit. But a lot of my colleagues put only one of them. He said after that he wanted X-linked, because the heredogram had more trends of X-linked diseases. I just don't bite that.
This is not the way we calculate probability. For example a autosomal disease can happen in 44 out of the 46 chromosomes while the X-linked only in the X chromossome (1.5 in average). Also, what if recessive diseases are more common than dominant. What if most of the people with X-linked diseases are infertile or if most of them dies. All of these reduces the chance of a given heredogram being Dominant X-linked. We can't say something one based on the trends.
The heredogram is the below.
Do you agree with me? I really need some opinions of other people so we can contest the teacher. Otherwise half of the classroom will get a zero.
Last edited by a moderator: