randombill
- 81
- 0
I tried to used the Crank-Nicolson method to solve the advection equation for a Gaussian pulse using the code on this page and I'm lost at trying to setup the initial values for the Gaussian which is passed as an array into the advection matrix.
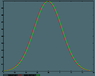
The Gaussian pulse is defined
initial condition: u(x,0) = exp[-100(x-0.5)^2] as on this page written for the Lax method.
The initial velocity is v = 1, timestep = 9.95x10^(-3), N = 200 and C = 2.
In the following code the matrix u is initialized to 202 elements and I try to initialize the matrix u
with the gaussian pulse for each index between 1 and 200 but something is very wrong. For one I'm not sure how to put the timestep into the gaussian equation. I suppose if x/t = v then tv = x and (9.95x10^(-3))*(velocity = 1) = distance then each timestep solved with advect1D would move the Gaussian to the right a small amount.
The thing I'm not sure about is this part in my code
I initialize every element of u(index) with the same gaussian? Well hopefully someone can help me here's the complete code.
The Gaussian pulse is defined
initial condition: u(x,0) = exp[-100(x-0.5)^2] as on this page written for the Lax method.
The initial velocity is v = 1, timestep = 9.95x10^(-3), N = 200 and C = 2.
In the following code the matrix u is initialized to 202 elements and I try to initialize the matrix u
with the gaussian pulse for each index between 1 and 200 but something is very wrong. For one I'm not sure how to put the timestep into the gaussian equation. I suppose if x/t = v then tv = x and (9.95x10^(-3))*(velocity = 1) = distance then each timestep solved with advect1D would move the Gaussian to the right a small amount.
The thing I'm not sure about is this part in my code
Code:
for (int index = 1; index < 201; index++)
{
u(index) = exp(-100*(distance - 0.5)*(distance-0.5));
}I initialize every element of u(index) with the same gaussian? Well hopefully someone can help me here's the complete code.
Code:
// Advect1D.cpp
// Function to evolve advection equation in 1-d:
// du / dt + v du / dx = 0 for xl <= x <= xh
// u = 0 at x=xl and x=xh
// Array u assumed to be of extent N+2.
// Now, ith element of array corresponds to
// x_i = xl + i * dx i=0,N+1
// Here, dx = (xh - xl) / (N+1) is grid spacing.
// Function evolves u by single time-step.
// C = v dt / dx, where dt is time-step.
// Uses Crank-Nicholson scheme.
#include <blitz/array.h>
#include <iostream>
#include <cmath>
using namespace blitz;
void Tridiagonal (Array<double,1> a, Array<double,1> b, Array<double,1> c, Array<double,1> w, Array<double,1>& u);
void Advect1D (Array<double,1>& u, double C);
int main()
{
Array<double,1> u(202);
u = 0;
double distance = 9.95e-3;
for (int index = 1; index < 201; index++)
{
u(index) = exp(-100*(distance - 0.5)*(distance-0.5));
}
double C = 2;
Advect1D ( u, C);
std::cout<<u<<std::endl;
return 0;
}
void Advect1D (Array<double,1>& u, double C)
{
// Find N. Declare local arrays.
int N = u.extent(0) - 2;
std::cout<<"N = "<<N<<std::endl;
Array<double,1> a(N+2), b(N+2), c(N+2), w(N+2);
a = 0;
b = 0;
c = 0;
w = 0;
// Initialize tridiagonal matrix
for (int i = 2; i <= N; i++)
a(i) = - 0.25 * C;
for (int i = 1; i <= N; i++)
b(i) = 1.;
for (int i = 1; i <= N-1; i++)
c(i) = + 0.25 * C;
// Initialize right-hand side vector
for (int i = 1; i <= N; i++)
w(i) = u(i) - 0.25 * C * (u(i+1) - u(i-1));
std::cout<<a<<std::endl;
std::cout<<b<<std::endl;
std::cout<<c<<std::endl;
std::cout<<w<<std::endl;
// Invert tridiagonal matrix equation
Tridiagonal (a, b, c, w, u);
// Calculate i=0 and i=N+1 values
u(0) = 0.;
u(N+1) = 0.;
}
void Tridiagonal (Array<double,1> a, Array<double,1> b, Array<double,1> c, Array<double,1> w, Array<double,1>& u)
{
// Find N. Declare local arrays.
int N = a.extent(0) - 2;
Array<double,1> x(N), y(N);
// Scan up diagonal from i = N to 1
x(N-1) = - a(N) / b(N);
y(N-1) = w(N) / b(N);
for (int i = N-2; i > 0; i--)
{
x(i) = - a(i+1) / (b(i+1) + c(i+1) * x(i+1));
y(i) = (w(i+1) - c(i+1) * y(i+1)) / (b(i+1) + c(i+1) * x(i+1));
}
x(0) = 0.;
y(0) = (w(1) - c(1) * y(1)) / (b(1) + c(1) * x(1));
// Scan down diagonal from i = 0 to N-1
u(1) = y(0);
for (int i = 1; i < N; i++)
u(i+1) = x(i) * u(i) + y(i);
}